|
As part of the strong NTU representation at the 2024 Annual Microbiology Conference, Dr Leanne Timpson and Mr Richard Perrotta’s abstract entitled “Building Alliances – An Interdisciplinary Approach to Learning about the Microscopic World” was selected for a Flash Talk at the Education and Outreach Symposium. The project was initiated in 2023 by Dr Leanne Timpson, Lecturer in Microbiology at Nottingham Trent University, who, during her own PhD in working with extremophilic microorganisms, often wondered about visualisation techniques for structures and molecular level processes that were too small to observe with the naked eye! Together with Mr Richard Perrotta, Lecturer in Art at the Confetti Institute of Creative Technologies, they created the “Building Alliances” interdisciplinary project between NTU and Confetti Institute. The collaborative project was possible as a result of Dr Timpson’s experience in working with microorganisms and Mr Perrotta’s experience in working as an inker and comic book artist for Marvel and DC comics. Both having experience in working in industrial and academic environments also supported the planning and running of this project. The “Building Alliances” project involved students from foundation-year Biosciences at Nottingham Trent University and second-year Games Art undergraduate students from Confetti Institute of Creative Technologies. Instead of bringing superheroes like Wolverine and Iron Man to life, this project involved using traditional drawing methods and digital artwork to peer inside the world of biology to include cells, microorganisms and microscopic structures. NTU Bioscience students developed professional identity and researching skills through teaching and providing editorial commentary on the work produced by the Confetti students. In turn, the Confetti students gained subject knowledge in an area they were previously unfamiliar with and developed their concept art drawing skills. Both cohorts developed essential work-like experience in terms of collaboration, communication, teamwork and compromise. Students realised the importance of the work-like element of the project in terms of meeting deadlines and working with others from disciplines and levels different to their own. One student commented “The ability to collaborate with people in very different fields of work and make something meaningful was great!” The work created was used by the Confetti students as part of their Games Art Toolkit portfolio. The Bioscience students also used the artwork as a resource when preparing for poster presentations on cells and their organelles. Both Leanne and Richard are extremely proud of what the students achieved during the first stage of this project. They are thankful to Dr Sarah Rayment and Dr Michael Loughlin at Nottingham Trent University for inspiring conversations and support with the ethics application as well as Mr Simon Holland at Confetti Institute of Creative Technologies for his support with this work.  Concept art produced from the “Building Alliances” project. Left-hand image: Artistic interpretation of a golgi apparatus. Credit: Anita Nagy (The Winning Swimmers/MICRO FROG GAMES). Middle image: Artistic interpretation of an Escherichia coli cell. Credit: Grace Haywood (Synaptic Knobs/Bionic Ink). Right-hand panel: Artistic interpretation of a deep-sea smoker (an example of an environment where thermophilic microorganism may be found). Credit: Will Forsyth (Biowizards/CELLSHADED STUDIOS).
0 Comments
The Microbiology Society holds one of the biggest conferences in the field annually. This year the conference was held in Edinburgh, between 8 and 11 April. Talks ranged across the microbial research landscape, with emphasis on viruses, antimicrobial resistance, microbiomes, genetics and education/outreach.
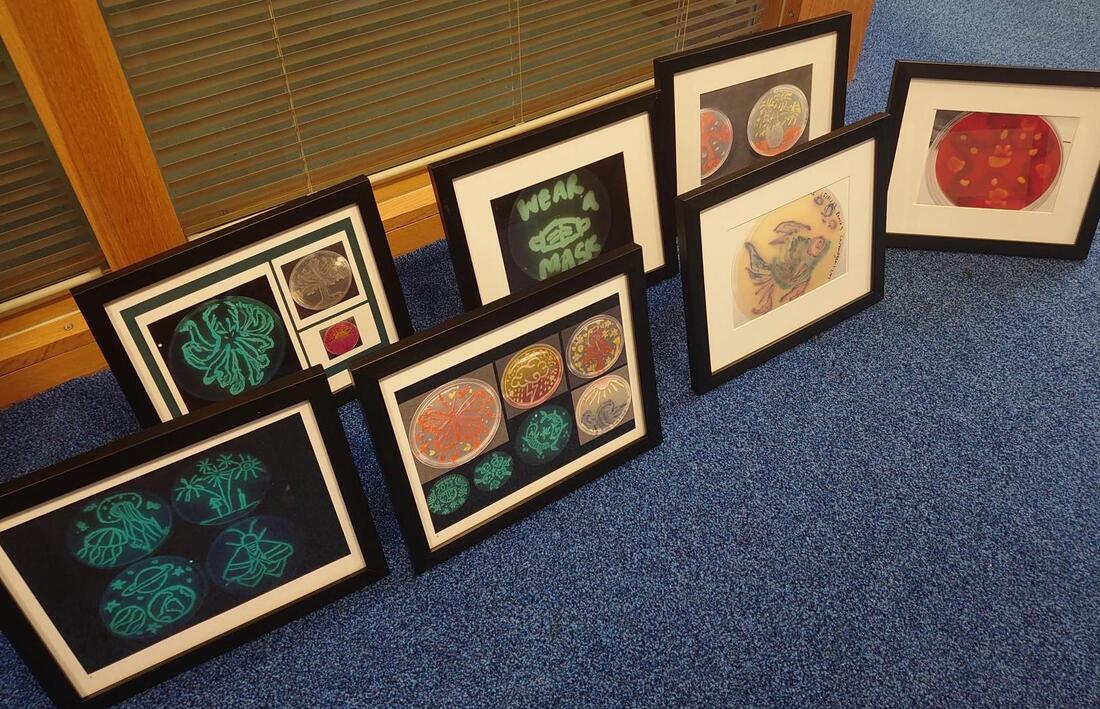
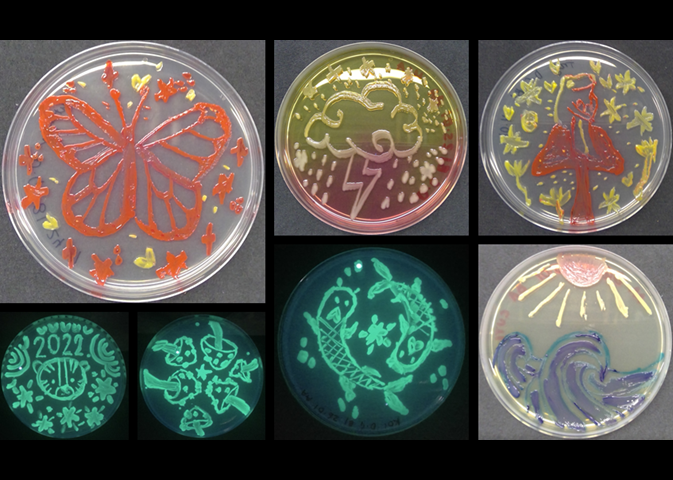
AROM was in strong attendance this year. Seventeen members (almost half!) of the group attended, with an almost even split between staff and students. We presented 17 posters ranging across topics from mechanisms and evolution of antimicrobial resistance to interactions between microbes and cancer cells, description of novel weberviruses, and various insights into computational methodologies. We also had a strong presence on our teaching and learning aspects, showing posters on the Bioskills-At-Home kits and interdisciplinary approaches to teaching microbiology. The latter was also selected for a flash talk. We gave another talk on isolation of Serratia bacteriophages as novel treatment approaches. Many AROM members of academic staff were also at the conference in their capacity as division members of boards or as editors for journals of the Microbiology Society. We thank the Department of Biosciences and the Centre for Systems Health and integrated Metabolic Research (SHiMR) for their support to attend the conference. It allowed for a lot of networking opportunities and showed the strong presence of microbiology at NTU. Our first agar art club ran for four consecutive evenings (24–27 January) with 34 students from our 2nd and 3rd year Biosciences' undergraduate courses. The students worked with pigmented bacteria, differential growth media and Escherichia coli expressing green fluorescent protein to produce a range of artistic effects, while also learning about media preparation, aseptic technique and the use of differential growth media for diagnostic purposes. The Head of Department's prize for outstanding artistic creations went to Danae Guerrero for her wonderful submissions. Aren’t they great? Well done to all participants, and congratulations to the competition winners! All our prize winners received framed photos of their wonderful creations. This event couldn’t have gone ahead without the support of our teaching technical team, led by David Wilkinson, and the following academics: Jody Winter, Karin Garrie, Lesley Hoyles and Maria Rosa Domingo-Sananes. A big thank you to all for putting together a hugely enjoyable range of activities.
We’re already looking forward to running similar events in the future! Gram-negative bacteria are defined by an accessory membrane, the outer membrane, covering the bacterial cell. This membrane has many unique features, such as it being an asymmetric bilayer where the outer leaflet is composed of lipopolysaccharide, a type of glycolipid exclusively found in Gram-negative bacteria. In addition, the integral membrane proteins of the outer membrane differ from those found in the cytoplasmic membrane of both prokaryotes and eukaryotes. The “standard” transmembrane proteins of the cytoplasmic membrane contain hydrophobic (fat-soluble) alpha-helices that anchor the proteins in the membrane. By contrast, the transmembrane proteins of the outer membrane have a different structure: they contain a beta-barrel domain, which could be considered to be a sheet-like structure where one face is hydrophobic and the other hydrophilic (or water-loving). The sheet closes in on itself to form a cylinder with a hydrophobic exterior and a hydrophilic interior. AROM’s Jack C. Leo and colleagues from Oslo have written a review focussing on the transmembrane beta-barrel proteins of Gram-negative bacteria. The review gives a general overview of these proteins, discussing their structure, folding, function, evolution as well as their medical relevance and role in antibiotic resistance.
Hermansen S, Linke D, Leo JC. Transmembrane beta-barrel proteins of bacteria: from structure to function. Adv Protein Chem Struct Biol. https://doi.org/10.1016/bs.apcsb.2021.07.002. Many important bacterial pathogens, including the important ESKAPE pathogen Klebsiella pneumoniae, encode and express polysaccharide capsules as their outermost cell structure. These structures are often key virulence factors, aiding in host immune evasion and providing protection from cellular and humoral immune responses. There is growing evidence that removal of these protective capsules, via administration of selective capsule depolymerases, results in rapid elimination of pathogens by the host. Such an approach should be successful against antibiotic-resistant isolates and, therefore, capsule depolymerases represent attractive future therapeutics.
In a recent publication, AROM’s David Negus and collaborators identified, cloned and tested a number of phage-derived capsule depolymerases selective for prevalent K. pneumoniae K-types (capsule types). In the study, over 60 lytic K. pneumoniae phages were isolated and analysis of whole-genome sequence data was used to identify phage-encoded depolymerases. Three of these enzymes were cloned and expressed as recombinant proteins. Using microscopy, the researchers show these depolymerases rapidly remove the capsule from the surface of K. pneumoniae isolates. These enzymes may provide the basis for a future therapy against antibiotic-resistant K. pneumoniae infections. Blundell-Hunter G, Enright MC, Negus D, et al. Characterisation of bacteriophage-encoded depolymerases selective for key Klebsiella pneumoniae capsular exopolysaccharides. Front Cell Infect Microbiol. 2021; doi.org/10.3389/fcimb.2021.686090. As part of developing an e-learning module on medical microbiology, AROM’s Jack Leo implemented new peer feedback activities in autumn 2020 to provide more opportunities for students to receive formative feedback on their coursework. The module material was mostly online, and the students were expected to go through the material independently. There were five 1-hour support sessions that were run online in autumn 2020. Apart from these support sessions, there were no opportunities for formative feedback in earlier years, and students did not receive any individual feedback. Therefore, Jack decided to implement the peer feedback activities for the 2020 module. For these, students were assigned to groups of four and they were encouraged to organise an hour’s meeting to give each other feedback on their coursework. Some written guidance was given in the module materials, but the activities were unsupervised and participation was voluntary.
As part of Jack’s academic apprenticeship in higher education, he undertook action research to evaluate the effect of the peer feedback activities on student satisfaction and learning. Of 54 students, 13 answered an online questionnaire on the feedback activities. of these, only one actually took part in a peer feedback activity. Therefore, the effect on student satisfaction could not be fully evaluated. However, the questionnaire did give insight into why most students did not participate. Many respondents cited a lack of time. This has been an issue this year, with the pandemic adding an extra layer of difficulty to what would already have been a busy year. Some students also noted that they felt uncomfortable with sharing their work. As students have not been able to meet in person, a lack of trust and cohesion within the cohort is understandable. Based on the online activity of the assigned groups, it seems that only a few students actually participated in the peer feedback activity. The one student who did take part and responded was largely positive towards the activities, and they found them to be useful. Some other students said they would have liked to take part, but other members in their group were not willing to meet. Several others noted that, in hindsight, taking part in the peer feedback activities might have been helpful. These findings could be useful for colleagues planning to implement similar kinds of unsupervised peer feedback activities. It seems that building trust within the cohort would be an important aspect to improve to promote participation. Jack will do this by including preliminary group activities (hopefully face to face rather than live) in future iterations of the module to allow the members to get to know each other before embarking on the extracurricular peer feedback activities. Beta-barrel outer membrane proteins of Gram-negative bacteria are inserted into the outer membrane by the BAM (beta-barrel assembly machinery) complex. In addition, many Gram-negative bacteria harbour a related complex, called the TAM (translocation and assembly machinery). The role of this second complex is not fully understood, but it is needed for the assembly of some autotransporters and pili. An international team of investigators from the UK, Germany, Norway and Australia, including AROM’s Jack Leo, investigated the requirement of the TAM and some BAM components for the assembly of trimeric autotransporter adhesins, a group of large, adhesive surface proteins that are involved in several diseases. The study shows that of the two complexes, only the essential core components of the BAM – BamA and BamD – are required for outer membrane incorporation and surface exposure of these proteins. Thus, these components might be targeted by novel anti-infective or antimicrobial compounds to prevent binding of bacteria to host cells.
Rooke JL, Icke C, Wells TJ, et al. BamA and BamD are essential for the secretion of trimeric autotransporter adhesins. Front Microbiol. 2020; doi.org/10.3389/fmicb.2021.628879. Enteropathogenic and enterohaemorrhagic Escherichia coli (EPEC and EHEC, respectively) are major causes of diarrhoeal disease. To cause disease, EPEC and EHEC must bind to the cells of the intestine. This causes rearrangements of the cell surface, resulting in a “pedestal” around the bacteria. The formation of this structure is necessary for intimate attachment of the bacteria to intestinal cells, which in turn is necessary for diarrhoea to develop.
Pedestal formation depends on a surface protein called intimin, which is an adhesin that mediates the intimate attachment of the pathogens to the intestinal cells. The adhesive tip of intimin is projected away from the bacterial surface by several tandemly arranged immunoglobulin (Ig)-like domains that form the “stalk” of the protein. Jack C. Leo and colleagues from Denmark, Norway, Sweden, Germany and Japan have solved the structure of two of the Ig-like domains, called D00 and D0, which are located at the beginning of the stalk. These have been the final missing pieces which allow full structural modelling of the extracellular region of intimin. The short connector between these two domains is rigid, whereas the connectors between the other Ig-like domains are more flexible. Using computer modelling, the researchers show that this arrangement of rigid and flexible regions increases the overall reach of the adhesin, i.e. allows it to project further on average from the bacterial surface than if all the connectors were flexible. The flexibility of some parts, on the other hand, could help the adhesin bind to its receptor on the host cell surface. Weikum J, Kulakova A, Tesei G., et al. The extracellular juncture domains in the intimin passenger adopt a constitutively extended conformation inducing restraints to its sphere of action. Sci Rep. 2020; doi.org/10.1038/s41598-020-77706-7. Dr Jody Winter recently contributed to a successful ERASMUS+ International Credit Mobility bid that will allow Microbiology staff and students from NTU and Makerere University, Uganda to take part in exchange visits to each other's universities. The project, funded for the next 3 years, will see NTU and Makerere students taking part in summer research placements in Uganda and the UK, respectively. This will be a unique opportunity for UK-based and African students to experience different educational environments and to work on projects that will contribute to strengthening research links between microbiologists at Makerere University and NTU.
The NTU Microbiology team, led by Dr Jody Winter, will also support joint research projects on plants with antimicrobial activities between NTU, Jomo Kenyatta University of Agriculture and Technology, Mount Kenya University and Brackenhurst Kenya Ltd as part of a second funded ERASMUS+ International Credit Mobility project. We are delighted to contribute to these projects and look forward to welcoming African microbiologists to NTU in the (hopefully) not-too-distant future. Antimicrobial stewardship promotes responsible use of antimicrobials to prevent the development of antimicrobial resistance. Implementation of antimicrobial stewardship remains poor in many low- and middle-income countries. Using a one-health approach, for the past 2 years a team of academics from Nottingham Trent University and Makerere University, Uganda and pharmacists and healthcare professionals from Buckinghamshire Healthcare NHS Trust have been working together on a project to strengthen antimicrobial stewardship in Wakiso District, Uganda. You can read all about the excellent outcomes from this exciting project here.
Musoke D, Kitutu FE, Mugisha L, et al. A one health approach to strengthening antimicrobial stewardship in Wakiso District, Uganda. Antibiotics. 2020; doi.org/10.3390/antibiotics9110764. This is not the end of NTU and AROM’s work on antimicrobial stewardship and resistance in Uganda. As part of a recently funded ERASMUS project, microbiology students from NTU and Makerere University will have the opportunity to work at each other’s universities on joint summer research placements. This will be a unique opportunity for UK-based and African students to experience different educational environments and to work on projects that will contribute to strengthening research links between Makerere University and NTU. The NTU Microbiology team, led by Dr Jody Winter, will also support joint research projects on plants with antimicrobial activities between NTU, Jomo Kenyatta University of Agriculture and Technology, Mount Kenya University and Brackenhurst Kenya Ltd as part of a second recently funded ERASMUS programme. |









 RSS Feed
RSS Feed
